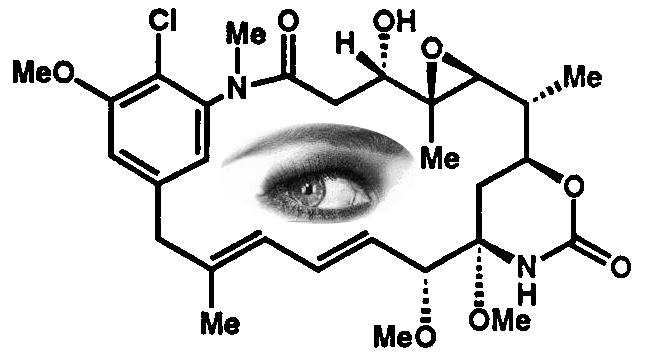
 Chemistry
is all about structure and, as readers of these pages well know, there are
countless packages for drawing and editing chemical structures. But, what
if you have a parcel of scanned reprint research papers or a pile of
downloads from which you would like to extract the chemical information?
One way would be to work your way through each paper, redrawing the
structures therein in one of those drawing packages. The problem, of
course, being that printed pictures, embedded gifs and the like carry with
them none of the underlying chemical information essential for creating an
accurate chemical structure with the atoms and bonds and the right places
that can be manipulated as a "real" chemical model albeit on the screen.
Chemistry
is all about structure and, as readers of these pages well know, there are
countless packages for drawing and editing chemical structures. But, what
if you have a parcel of scanned reprint research papers or a pile of
downloads from which you would like to extract the chemical information?
One way would be to work your way through each paper, redrawing the
structures therein in one of those drawing packages. The problem, of
course, being that printed pictures, embedded gifs and the like carry with
them none of the underlying chemical information essential for creating an
accurate chemical structure with the atoms and bonds and the right places
that can be manipulated as a "real" chemical model albeit on the screen.Enter Clide.
Clide - Chemical Literature Data Extraction project - does for chemists and drug discovery researchers what OCR (optical character recognition) does for wordsmiths. It turns flat two-dimensional semantics free representations and turns them into the "real" thing. Take a bitmap image or an Adobe portable document format (PDF) file of that molecular structure, run it through Clide and the product is the more familiar, but crucially manipulable, chemical structure.
The program started out life as a project in the Department of Chemistry at the University of Leeds, UK, and is now a commercially available entity that comes in two flavours. A full version and a Lite version. Clide extracts information from chemical literature then store it in a database. The internal representation of Clide stores ALL information included in the structure in such a way that it can be converted to other formats. The input of the program is a bitmap image (BMP) of a chemical document or PDF file.
 The program outputs information as an ASCII file containing the recognised
structures, the reactions and any text associated with the image. This is
stored in the Clide database. By default, information is held in Clide
file format, which contains all the information that has been processed by
Clide. However, the information can be exported in other formats such as
the commonly used MDL molfile and CambridgeSoft's ChemDraw v 6.0 format,
mac ChemDraw is also available as are two formatting and printing systems
Postscript and text only. This makes Clide a rather versatile character is
these formats can be handled by more sophisticated chemical spreadsheets
and databases. They also allow the extracted structures to be easily
edited, displayed and converted in a variety of useful ways. A mol file,
for instance, can be rendered in three dimensions on screen using MDL's
associated Chime program, or its precursor system Rasmol, in a web
browser). However, these proprietary formats contain only the raw
molecular structure information that their design allows. The molfile
contains the connection tables of the structures, for example, so page
numbers, titles and associated text and images are lost in converting to
this format.
The program outputs information as an ASCII file containing the recognised
structures, the reactions and any text associated with the image. This is
stored in the Clide database. By default, information is held in Clide
file format, which contains all the information that has been processed by
Clide. However, the information can be exported in other formats such as
the commonly used MDL molfile and CambridgeSoft's ChemDraw v 6.0 format,
mac ChemDraw is also available as are two formatting and printing systems
Postscript and text only. This makes Clide a rather versatile character is
these formats can be handled by more sophisticated chemical spreadsheets
and databases. They also allow the extracted structures to be easily
edited, displayed and converted in a variety of useful ways. A mol file,
for instance, can be rendered in three dimensions on screen using MDL's
associated Chime program, or its precursor system Rasmol, in a web
browser). However, these proprietary formats contain only the raw
molecular structure information that their design allows. The molfile
contains the connection tables of the structures, for example, so page
numbers, titles and associated text and images are lost in converting to
this format.Clide document/image processing takes place in three constituent parts. First, is the physical document structure analysis. This consists of the identification of the connected components of the image, what Clide makes by loading an image and the demarcation of the image into its graphic and textual regions. Bibliographical information (i.e. title and author of a document) is extracted automatically.
Second is the recognition of the primitives. This starts with classification of the connected components into characters, lines and graphics. This is followed by the application of the recognition process to these elements by Clide's sophisticated algorithms. The characters are recognised by the program's OCR module while the graphic lines are recognised by the graphic recognition module.
And, finally, the logical document structure recognition. The page is analysed and chemical structures recognised as logical units. This involves building the connection table of the structure that relates each point and line to an atom and a bond. At this stage, reactions too are recognised so that reactant and product can be differentiated. Generic text is also identified at this stage and checked, or parsed. Lastly, any reaction text is then identified and parsed so that it is not lost into the milieu of the body text but rather retained within any reaction scheme.
The system can also extract generic structure information from graphics and text and either expand or condense such structures before they are entered into a database. The whole process takes seconds per "printed" page on the meanest of PCs (e.g. a 366 MHz PII laptop with 160 MB memory takes less than half a second to process a bitmap image to MDL Molfile format).
It is a fantastic program in concept and I can see it being developed much further for large pharmaceutical company databases and abstracting services once the programmers have ironed out the residual bugs. Clide could also dovetail nicely with Simbiosys' simulated biomolecular systems in which the company provides a software system for rational drug design and the optimization of small, organic therapeutic molecules. One additional area in which Clide could lend a helping hand is in the creation of a truly chemically literate web search engine.
Although the application of chemoinformatics in the drug discovery process has drastically grown in the past two decades, to the point that it is currently being applied at all stages of the process, there is still huge potential for growth. Integrated data storage and retrieval solutions, that interconnect and provide feedback to the various links in the chain are increasingly desirable. Chemically intelligent solutions like Clide will help produce successful drugs faster and with less cost.
Clide is available for the MS Windows 2000/NT 4.0/XP/Me/98/95 platforms.