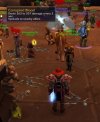
In September 2005, about 4 million World of Warcraft gamers saw a new and unexpected challenge in the game. Players exploring a new area within the game encountered an extremely virulent, highly contagious disease, known as Corrupted Blood, which had been introduced in an update on the 13th of that month. The disease quickly spread, like the Black Death, to the population centres of the fantasy world, killing many and causing social chaos. But, who, other than the gamers themselves should care about virtual deaths and digital disease?
Well, according to Eric Lofgren of Tufts University School of Medicine, in Boston and Nina Fefferman of Rutgers University, in Piscataway, we all should care because the virtual epidemic could provide a very useful model of how diseases spread, how individuals and groups respond to the presence of a killer disease, and what we might do to control an outbreak, of bird flu or a SARS-type disease, in the real world. Writing in the journal Lancet Infectious Diseases, the team explains how simulation models are very useful in studying the spread of disease, epidemiology. However, validating the models or tailoring them to particular human behaviour patterns is next to impossible. The World of Warcraft incident, known as the Corrupted Blood outbreak, on the other hand, “provided an excellent example of the potential of such systems.” They explain that while data from the Corrupted Blood outbreak were not gathered scientifically at the time and so represent a missed opportunity, there is the potential for deliberately engineering such gameplay into other virtual worlds. “Virtual outbreaks designed and implemented with public-health studies in mind have the potential to bridge the gap between traditional epidemiological studies on populations and large-scale computer simulations,” the researchers say, “these would involve both unprogrammed human behaviour and large numbers of test participants in a controlled environment where the disease parameters are known.”
The use of distributed networks to help solve scientific and medical problems is not new. many Internet users will have heard of the likes of SETI@home. This system is essentially a program you install on your computer which uses idle time to search for signs of extra terrestrial life in downloaded astronomical data.
SETI@home is just one of a group of applications based on BOINC, the Berkeley Open Infrastructure for Network Computing. You can use your Windows, Mac, or Linux machine to help find cures for disease, study climate change, discover pulsars and solve various other problems in earth sciences, astronomy, physics, biology, medicine, mathematics and strategy games. You can find a full list of projects here.
A recent paper in the International Journal of Web and Grid Services (2007, 3, pp 354-368) reviewed the state of the art in such distributed applications the world over. According to Bertil Schmidt of the University of New South Wales, Australia, desktop grid computing, as these kinds of distributed applications are known technically, is a relatively new technology but can nevertheless provide massive computing power for a variety of applications. He points out that the cost is low for the researchers, given that once set loose in the wild, the running costs of the program are themselves distributed to the downloaders’ computer and the costs to the researchers are then only in terms of retrieving results from those machines as and when necessary and analysing the incoming data.
Schmidt explains how BOINC provides “a proven open-source infrastructure to set up such projects in a relatively short time” and surveys the scientific projects, e-science, that have adopted this strategy. “The power and mass appeal of desktop grid computing for implementing task-parallel problems have been demonstrated in projects such as SETI@home,” Schmidt explains, pointing out that as of April 2007, the average performance of SETI@home was around 250 teraflops. A teraflops (sometimes teraFLOPS), is a million million floating point operations (or instructions) per second. The combined average performance of all BOINC-based projects was around 0.5 petaflops spread over more than 400 000 active CPUs, which is more powerful in total than IBM’s BlueGene/L, which peaks at 0.280 petaFLOPS. By comparison, the next generation supercomputer, Blue Gene/P, will run at 3000 teraflops, or 3 petaFLOPS, so this distributed power represents a vast resource for e-science and is as yet only very partially tapped.
Personally, I have the World Community Grid running on my computer. This application allows you to help out in the fight against Dengue fever, AIDS, and phase 2 of the human proteome folding project. At the time of writing, teh project has 318,888 members and is running on 715,025 devices amounting to a total of 107,837 years of computing time so far. You can view the latest stats here.
If you have a fast computer and are not running CPU or memory intensive applications, then you could do even more with any of those e-science projects, allowing the BOINC application to run even when your machine is not idle. You will not only accrue more “credits and kudos” on your chosen project but you could just solve one of the problems facing humanity.